Final ID: TU190
Comparison of the Olink and Alamar Paired Antibody-based Plasma Proteomics: The Dallas Heart Study
Abstract Body: Introduction Plasma proteomics provide valuable insights into circulating biomarkers of disease risk associated molecular pathways. However, challenges persist for low abundance proteins which are often biologically active. Newer assays, such as the Alamar NULISA assay, have improved sensitivity, but data regarding their performance characteristics relative to established large-scale assays such as Olink are limited.
Methods Among 179 participants in the multi-ethnic and community-based Dallas Heart Study who participated in the third study phase (2020-2024), we measured proteomics using two paired antibody-based methods (Alamar Biosciences inflammation panel; Olink Explore HT). For proteins represented in both datasets with values above limit of detection in at least half the population, we calculated cross-platform Spearman correlations. Blind duplicates were run for 9 samples and data was used to calculate coefficient of variation (CV) for each protein in both datasets. Finally, we tested associations of protein level with HF case/control status in a subset of 118 participants (59 paired case/control samples) using linear regression models adjusted for age, sex, and eGFR. The Benjamini-Hochberg method was used for multiple testing correction.
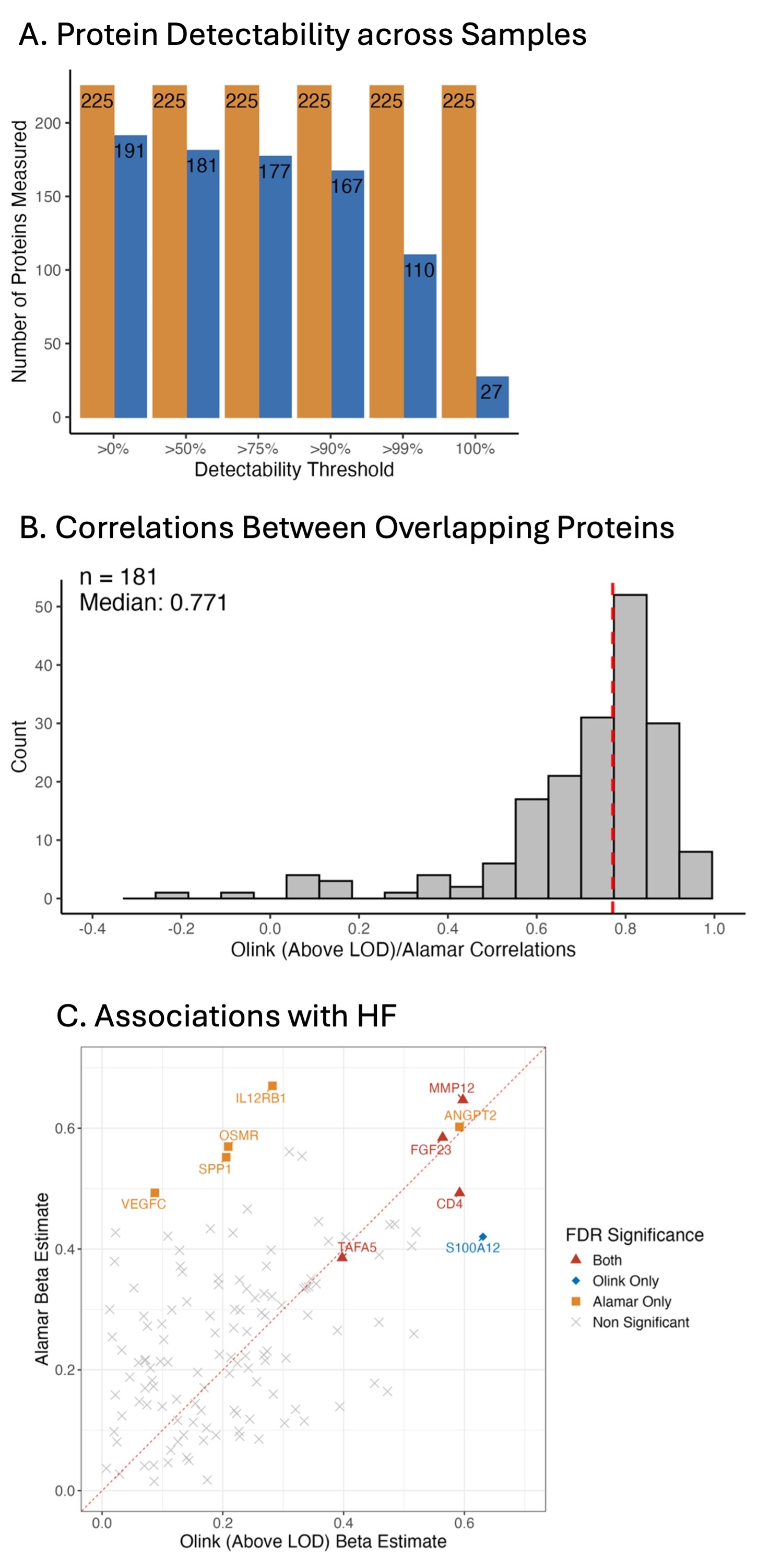
Results Mean age was 64±10 years, 59% were female, and 76% reported Black race. There were 248 proteins measured by Alamar and 2,352 proteins measured by Olink, of which 225 were captured by both platforms. Of these 225 proteins, all were detectable in >50% of samples by Alamar and 181 by Olink (Figure A). The median inter-assay correlation for these 181 proteins was 0.77 (Figure B). Assay variability was similar for both platforms (CV 7.5% for Olink, 6.9% for Alamar). After multiple testing correction, 9 proteins were associated with HF case/control status using Alamar data and 5 using Olink data (FDR adjusted p < 0.05; Figure C).
Conclusions Protein values from these two paired antibody-based proteomics methods are highly correlated, with the Alamar panel demonstrating greater protein detectability across samples despite similar assay variability. The increased sensitivity should be weighed against the decrease in protein coverage when deciding which platform to use for analyses.
Methods Among 179 participants in the multi-ethnic and community-based Dallas Heart Study who participated in the third study phase (2020-2024), we measured proteomics using two paired antibody-based methods (Alamar Biosciences inflammation panel; Olink Explore HT). For proteins represented in both datasets with values above limit of detection in at least half the population, we calculated cross-platform Spearman correlations. Blind duplicates were run for 9 samples and data was used to calculate coefficient of variation (CV) for each protein in both datasets. Finally, we tested associations of protein level with HF case/control status in a subset of 118 participants (59 paired case/control samples) using linear regression models adjusted for age, sex, and eGFR. The Benjamini-Hochberg method was used for multiple testing correction.
Results Mean age was 64±10 years, 59% were female, and 76% reported Black race. There were 248 proteins measured by Alamar and 2,352 proteins measured by Olink, of which 225 were captured by both platforms. Of these 225 proteins, all were detectable in >50% of samples by Alamar and 181 by Olink (Figure A). The median inter-assay correlation for these 181 proteins was 0.77 (Figure B). Assay variability was similar for both platforms (CV 7.5% for Olink, 6.9% for Alamar). After multiple testing correction, 9 proteins were associated with HF case/control status using Alamar data and 5 using Olink data (FDR adjusted p < 0.05; Figure C).
Conclusions Protein values from these two paired antibody-based proteomics methods are highly correlated, with the Alamar panel demonstrating greater protein detectability across samples despite similar assay variability. The increased sensitivity should be weighed against the decrease in protein coverage when deciding which platform to use for analyses.
More abstracts on this topic:
Acetylation of Electron Transfer Flavoprotein Alpha Is a Possible Regulatory Mechanism of Fatty Acid Oxidation in Diabetic Hearts
Tatekoshi Yuki, Yano Masaki, Hosoda Ryusuke, Saga Yukika, Kuno Atsushi
Anti-inflammatory effects of colchicine after myocardial infarctionMuroke Valtteri, Diaz Rafael, Maggioni Aldo Pietro, Pinto Fausto, Kouz Simon, Berry Colin, Lopez-sendon Jose, Koenig Wolfgang, Cossette Marieve, Guertin Marie-claude, Rheaume Eric, Rhainds David, Tardif Jean-claude, Brodeur Mathieu, Marchand Claude, Charpentier Daniel, Pedneault-gagnon Valerie, Rivest Patricia, Latour Frederic, Roubille Francois