Final ID: 04
Proteomics insights into heterogeneous associations between adiposity measures and all-cause mortality
Abstract Body: Background: Heterogeneity of associations between adiposity traits and all-cause mortality is well-reported, but its modifiable contributors are poorly characterized. Proteomics systematically characterizes physiology and pathophysiology of health. No study has comprehensively investigated effect-measure modifications of proteomics on the mortality risk associated with adiposity traits.
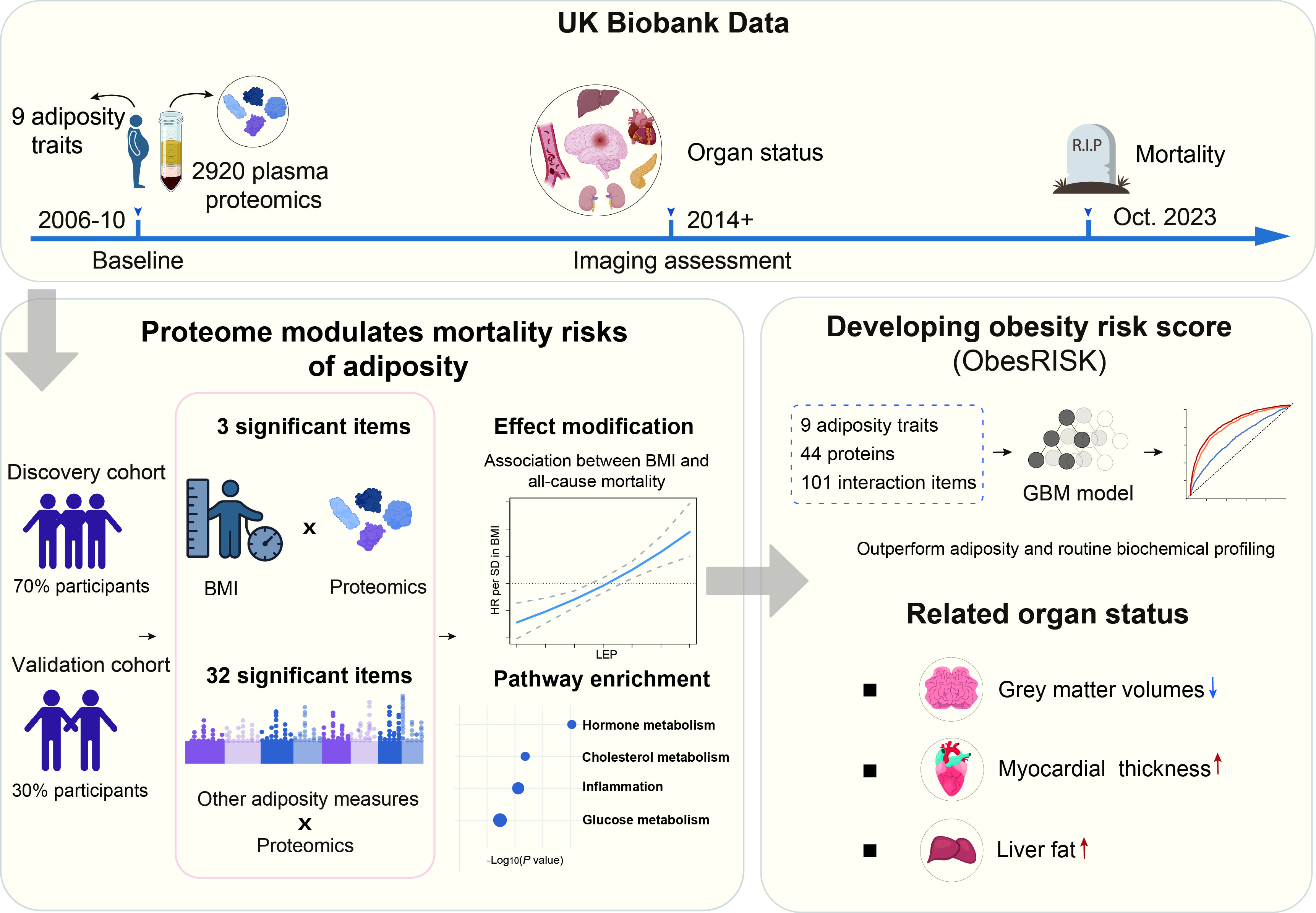
Methods: Using the UK Biobank data, we evaluated effect-measure modifications of proteomics on all-cause mortality risk associated with nine adiposity traits (i.e., BMI, waist circumference, waist-to-hip ratio, hip circumference, whole body fat mass, arm fat mass, leg fat mass, trunk fat mass, and VAT) using Cox regressions. Restricted cubic splines were implemented to model effect-measure modifications for proteins on the associations between adiposity and all-cause mortality. Through integrating adiposity, proteins, and interaction terms, we developed ObesRISK for predicting all-cause mortality risk using gradient boosting machine. Discrimination was assessed using Harrell C-statistics. We further estimated associations of ObesRISK with multi-organ traits using multivariable linear models.
Results: This study analyzed proteomics data from 46,233 participants. During a median follow-up of 14.6 years, 4621 deaths were documented. After applying Bonferroni correction, ADIPOQ (Pint=4.1×10-6), LEP (Pint=7.8×10-10), and RTN4R (Pint=3.9×10-7) modified the associations between BMI and all-cause mortality. We further identified 16 proteins that modified the all-cause mortality risk associated with the other eight adiposity traits. Pathway analyses mapped these proteins to inflammation, hormones (estradiol and glucocorticoid), and lipid metabolism. The mortality risk of adiposity varied depending on different levels of the identified proteins (all Bonferroni-corrected Pint<0.05). Empowered by protein-adiposity interactions, ObesRISK outperformed adiposity and routine biochemical profiling in predicting all-cause mortality (ΔC-index=0.036, 95%CI: 0.027-0.042). Furthermore, ObesRISK was associated with multi-organ dysfunctions including the brain, cardiac, and metabolic systems.
Conclusion: This study advances the understanding of heterogeneities in mortality risk associated with adiposity traits by utilizing a proteome-wide interaction modelling framework, thereby providing a proteomics approach that may improve precision risk classification and risk prediction of all-cause mortality.
Methods: Using the UK Biobank data, we evaluated effect-measure modifications of proteomics on all-cause mortality risk associated with nine adiposity traits (i.e., BMI, waist circumference, waist-to-hip ratio, hip circumference, whole body fat mass, arm fat mass, leg fat mass, trunk fat mass, and VAT) using Cox regressions. Restricted cubic splines were implemented to model effect-measure modifications for proteins on the associations between adiposity and all-cause mortality. Through integrating adiposity, proteins, and interaction terms, we developed ObesRISK for predicting all-cause mortality risk using gradient boosting machine. Discrimination was assessed using Harrell C-statistics. We further estimated associations of ObesRISK with multi-organ traits using multivariable linear models.
Results: This study analyzed proteomics data from 46,233 participants. During a median follow-up of 14.6 years, 4621 deaths were documented. After applying Bonferroni correction, ADIPOQ (Pint=4.1×10-6), LEP (Pint=7.8×10-10), and RTN4R (Pint=3.9×10-7) modified the associations between BMI and all-cause mortality. We further identified 16 proteins that modified the all-cause mortality risk associated with the other eight adiposity traits. Pathway analyses mapped these proteins to inflammation, hormones (estradiol and glucocorticoid), and lipid metabolism. The mortality risk of adiposity varied depending on different levels of the identified proteins (all Bonferroni-corrected Pint<0.05). Empowered by protein-adiposity interactions, ObesRISK outperformed adiposity and routine biochemical profiling in predicting all-cause mortality (ΔC-index=0.036, 95%CI: 0.027-0.042). Furthermore, ObesRISK was associated with multi-organ dysfunctions including the brain, cardiac, and metabolic systems.
Conclusion: This study advances the understanding of heterogeneities in mortality risk associated with adiposity traits by utilizing a proteome-wide interaction modelling framework, thereby providing a proteomics approach that may improve precision risk classification and risk prediction of all-cause mortality.
More abstracts on this topic:
Accelerometer-Measured Physical Activity Improves Predictive Validity of Fried Frailty Phenotype for All-Cause and Cardiovascular Disease Mortality: UK Biobank
Kong Lingsong, Kim Dae Hyun, Lee Chi Hyun, Spracklen Cassandra, Sturgeon Susan, Sirard John, Paluch Amanda
Apolipoprotein A-I Proteoforms, Cardiometabolic Status, and Coronary Heart Disease: Insights from the Dallas Heart StudyGangwar Anamika, Des Soye Benjamin, Saldanha Suzanne, Jaiswal Shailesh, Patel Parthvi Bharatkumar, Shah Amil, Pandey Ambarish, Wilkins John, Rohatgi Anand