Final ID: Sa3079
Cannabidiol alleviates fibrosis through AGE-RAGE axis inhibition in a Non-Ischemic Mouse Model of Heart Failure
Abstract Body (Do not enter title and authors here): Background
Cannabidiol has been shown recently to improve cardiac function in mouse models of heart failure with a potential mechanism through influencing mitochondrial bioenergetics. Specific mechanisms of this beneficial impact have not been clearly elucidated. In this study, we aimed to utilize gene expression data to study Cannabidiol’s benefit in a murine model of non-ischemic heart failure (HF).
Hypothesis
Cannabidiol ameliorates HF through genetic regulation of cardiometabolic pathways.
Methods
HF was induced in 3-month-old male C57BL/6 mice by administering drinking water containing 1% NaCl and 0.3 mg/mL L-NAME for one week. Subsequently, subcutaneous osmotic mini pumps were implanted to deliver angiotensin II (0.7 mg/kg/day) for 4 weeks. Pharmaceutical grade Cannabidiol was administered subcutaneously twice weekly (1 mg/kg). At week 5, myocardial tissue was harvested for transcriptomics. Bulk RNA sequencing was performed on cardiac tissue (3 mice per group) using an Illumina HiSeq 4000 platform. Paired-end reads were assembled and counted with StringTie. Differential expression analysis was performed with DESeq2. Gene set enrichment analysis (GSEA) was done with gseKEGG of the clusterProfiler 4.0. Visualization was carried out with enrichplot and GOPlot in R version 4.2.2.
Results
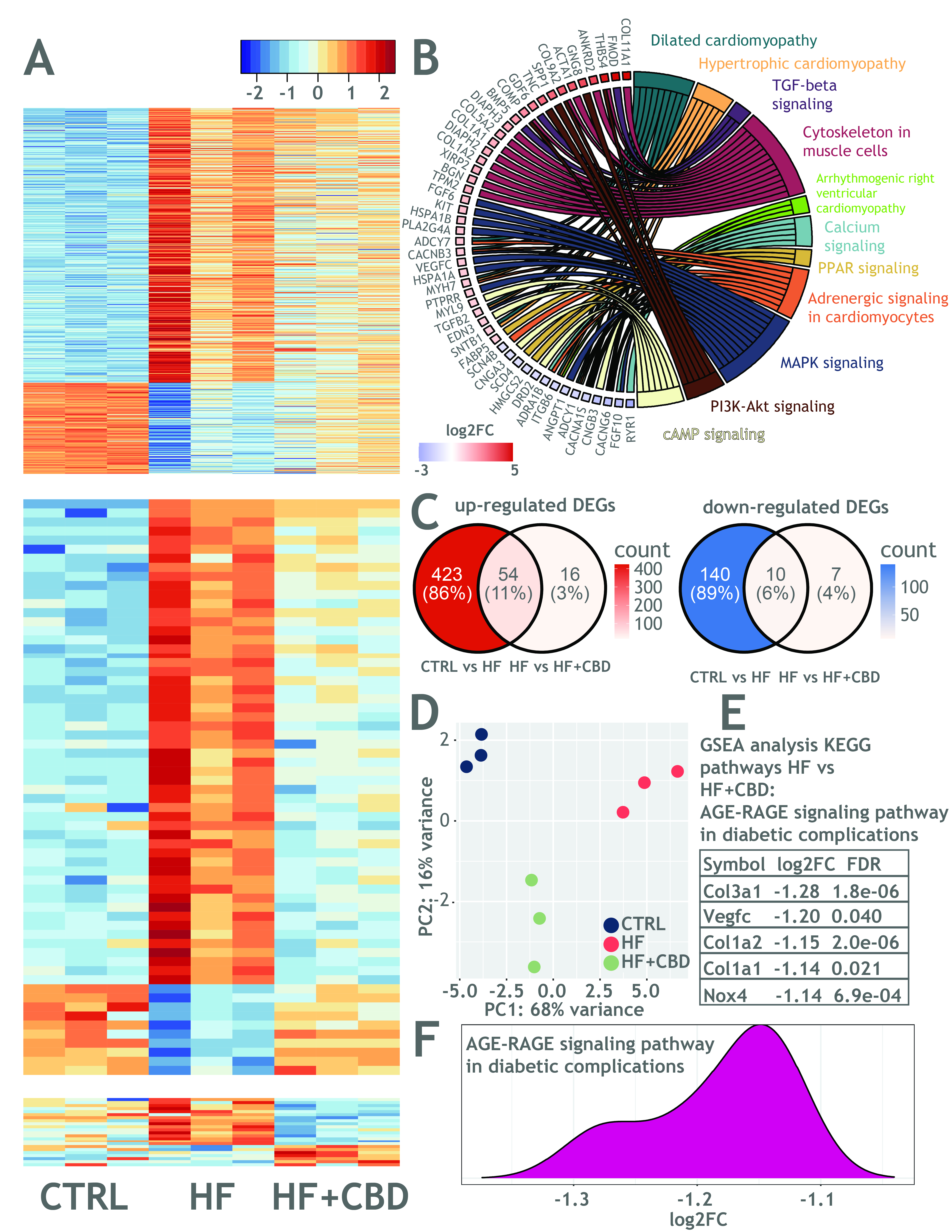
Relative to controls (CTRL), the heart of HF animals exhibited 627 differentially expressed genes (DEGs) (Fig1A top), involved in pathways such as dilated and hypertrophic cardiomyopathy, TGFβ signaling, cytoskeleton in muscle cells (chord plot in Fig1B), confirming the HF remodeling response. Intriguingly, 54 of the up-regulated and 10 of the down-regulated DEGs went back to CTRL expression levels after Cannabidiol administration (Fig1A middle, Fig1C). Overall, the transcriptomic profile of HF+ Cannabidiol animals was closer to the CTRL than to HF group, as evidenced by PCA analysis (Fig1D). Notably, twelve collagen-encoding genes were down-regulated in HF+ Cannabidiol relative to HF. GSEA revealed the AGE-RAGE pathway to be enriched in HF+ Cannabidiol vs. HF. DEGs contained in the AGE-RAGE pathway displayed negative FCs (Fig1E, F), possibly meaning a Cannabidiol-driven inhibition of the AGE-RAGE axis, a pathway known to be involved in chronic inflammation and aging.
Conclusion
Cannabidiol down-regulated genes related to extracellular matrix collagen fibers and the AGE-RAGE axis, possibly related to the alleviated fibrosis and inflammation in the HF+ Cannabidiol mice.
Cannabidiol has been shown recently to improve cardiac function in mouse models of heart failure with a potential mechanism through influencing mitochondrial bioenergetics. Specific mechanisms of this beneficial impact have not been clearly elucidated. In this study, we aimed to utilize gene expression data to study Cannabidiol’s benefit in a murine model of non-ischemic heart failure (HF).
Hypothesis
Cannabidiol ameliorates HF through genetic regulation of cardiometabolic pathways.
Methods
HF was induced in 3-month-old male C57BL/6 mice by administering drinking water containing 1% NaCl and 0.3 mg/mL L-NAME for one week. Subsequently, subcutaneous osmotic mini pumps were implanted to deliver angiotensin II (0.7 mg/kg/day) for 4 weeks. Pharmaceutical grade Cannabidiol was administered subcutaneously twice weekly (1 mg/kg). At week 5, myocardial tissue was harvested for transcriptomics. Bulk RNA sequencing was performed on cardiac tissue (3 mice per group) using an Illumina HiSeq 4000 platform. Paired-end reads were assembled and counted with StringTie. Differential expression analysis was performed with DESeq2. Gene set enrichment analysis (GSEA) was done with gseKEGG of the clusterProfiler 4.0. Visualization was carried out with enrichplot and GOPlot in R version 4.2.2.
Results
Relative to controls (CTRL), the heart of HF animals exhibited 627 differentially expressed genes (DEGs) (Fig1A top), involved in pathways such as dilated and hypertrophic cardiomyopathy, TGFβ signaling, cytoskeleton in muscle cells (chord plot in Fig1B), confirming the HF remodeling response. Intriguingly, 54 of the up-regulated and 10 of the down-regulated DEGs went back to CTRL expression levels after Cannabidiol administration (Fig1A middle, Fig1C). Overall, the transcriptomic profile of HF+ Cannabidiol animals was closer to the CTRL than to HF group, as evidenced by PCA analysis (Fig1D). Notably, twelve collagen-encoding genes were down-regulated in HF+ Cannabidiol relative to HF. GSEA revealed the AGE-RAGE pathway to be enriched in HF+ Cannabidiol vs. HF. DEGs contained in the AGE-RAGE pathway displayed negative FCs (Fig1E, F), possibly meaning a Cannabidiol-driven inhibition of the AGE-RAGE axis, a pathway known to be involved in chronic inflammation and aging.
Conclusion
Cannabidiol down-regulated genes related to extracellular matrix collagen fibers and the AGE-RAGE axis, possibly related to the alleviated fibrosis and inflammation in the HF+ Cannabidiol mice.
More abstracts on this topic:
A new mouse model for the study of cardiogenic TFs Tbx5, Gata4, and Mef2c during cardiogenesis
Leonard Riley, Sweat Mason, Eliason Steve, Zimmerman Kathy, Zhao Yi, Li Xiao, Hatcher Cathy, Weiss Robert, Amendt Brad
Ablation of Nephron Tubule Cell-Specific Npr1 Triggers High Blood Pressure and DysfunctionNeelamegam Kandasamy, Ramasamy Chandramohan, Pandey Kailash